Key take-aways
- LLM-driven evolution is an efficient, general method for code and agent optimization.
- We’ve already used code evolution in the development of Vet, our coding agent verifier, and to more than double a model’s reasoning performance in ARC-AGI tasks.
- Today, we are open-sourcing the Darwinian Evolver tool.
Introduction
Imagine you’re building an agentic, LLM-based application. You’ve gotten your hands on a couple of data points to evaluate your system’s performance, including some for which your application doesn’t quite work yet.
Unfortunately, you quickly find that it’s not always obvious how to improve the performance of your system. You tweak a prompt to fix one issue, just to find that your change causes a regression on the other data points. You might fiddle with the tools and chaining logic that wraps your LLM calls, just to find that your previous prompt no longer works with the new tools. In general, optimizing an LLM-based system end-to-end oftentimes turns into a highly manual and tedious process.
Wouldn’t it be nice if you could just ask a computer to optimize your software for you?
It was exactly this type of problem that we tried to solve during the development of our agent verifier Vet (Verify Everything). Vet uses LLMs and agents under the hood to verify the work of a separate coding agent and suggest further improvements. Existing prompt optimization frameworks, such as DSPy’s MIPRO, did not work for our use case. The existing techniques relied heavily on few-shot prompting, which wasn’t feasible in our case due to context length constraints. More fundamentally, the existing tools were limited to optimizing only a single prompt in isolation, without simultaneously considering the harnesses and decision logic surrounding it.
To solve this challenge, we turned to open-ended, evolution-based methods. Inspired by recent success from Sakana.ai with their Darwin Gödel Machines, we developed an in-house code evolution tool that could optimize Vet end-to-end.
To our surprise, we found that the exact same approach also yielded state-of-the-art results across a wide range of different optimization tasks. When we used our evolver to solve ARC-AGI tasks, we found that it was also very effective at reasoning, far exceeding the capabilities of the underlying base model. You can read about how we applied evolution to achieve SotA results in ARC-AGI-2 in our separate post.
LLM-based evolution is a near-universal optimizer
Our Darwinian Evolver provides an highly universal type of optimizer: An optimizer that can operate on near arbitrary code and text problems, and which does not require the solution space nor the scoring function to be differentiable.
Any problem for which potential solutions can a) be understood and modified by an LLM, and b) for which the quality of such a solution can be scored at least approximately, is in principle suitable for being optimized by our evolver.
This flexibility makes the evolutionary framework especially suitable for code and prompt optimization - both domains that behave highly non-linearly and are not inherently differentiable. Furthermore, the evolution-based approach is robust against the non-determinism involved in evaluating LLM-based systems, and is able to escape local optimization maxima thanks to its stochastic properties.
Notably, the evolution process is, in principle, open ended. There is no inherent limit to how much it can improve a given starting solution, as long as it is given sufficient time to do so.
In practice, the scoring data set and scoring methodology can impose a performance ceiling if they are too easily saturated. The strength of the mutator LLM (more about that later) can also impose a ceiling, though we’ve found empirically that our implementation is able to exceed the base model’s one-shot strength by a wide margin.
How to evolve code
Our high-level approach is as follows: We maintain a population of “organisms”, typically a piece of code. Starting with an initial organism, we repeatedly sample a parent organism and apply one or more mutators to generate children. These children get scored and then are added back to the population to be sampled in future iterations.
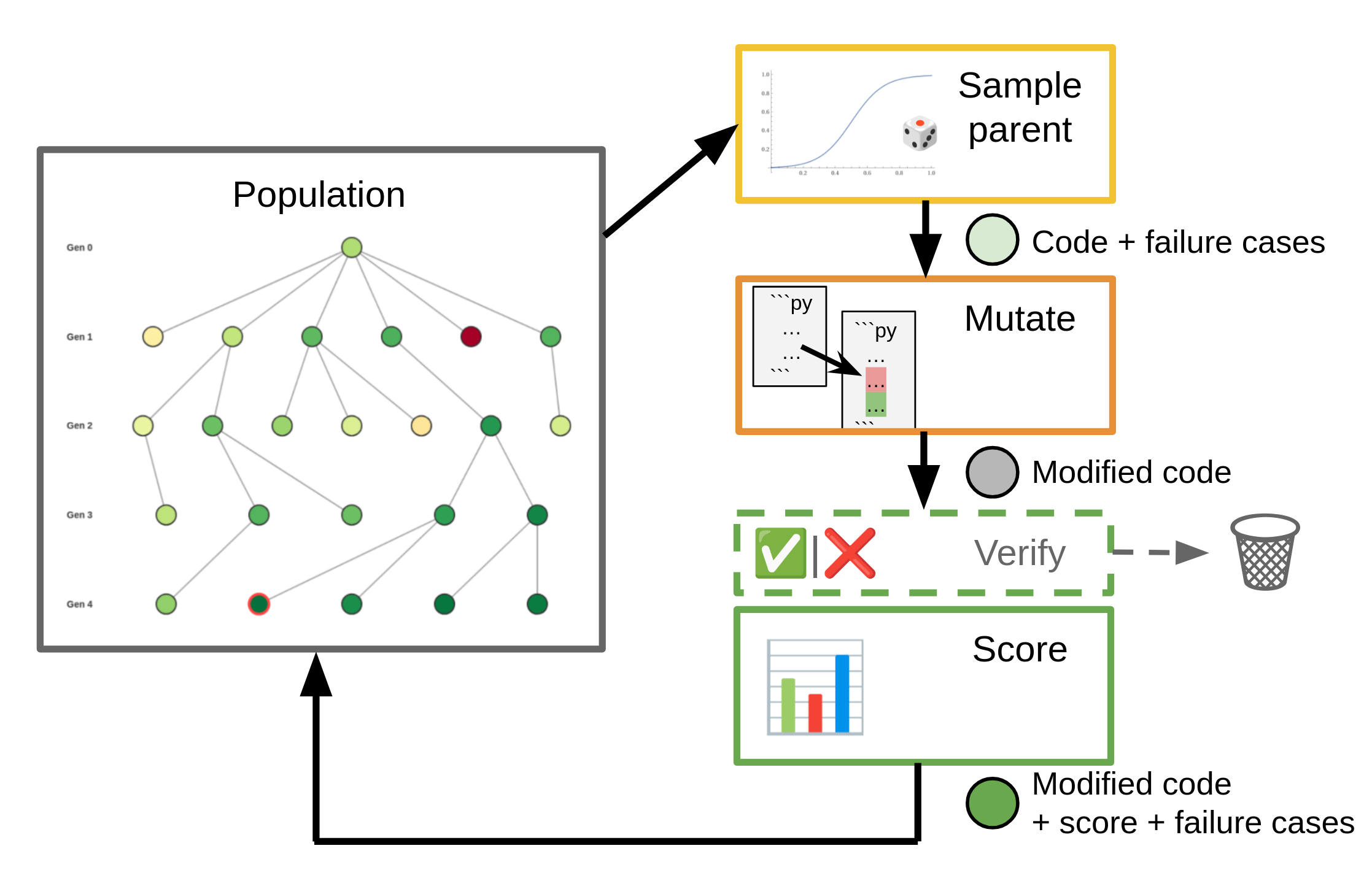
Our Darwinian Evolver is inspired by Sakana.ai’s Darwin Gödel Machines and their subsequent ShinkaEvolve framework, but incorporates several of our own enhancements and refinements. Darwin Gödel Machines were originally designed around the idea of self-improving coding agents. However, it turns out that the evolutionary framework behind them can be applied to a large range of coding and optimization problems.
Scoring fitness
There are several ways to define a fitness score for code, and the exact scoring approach will depend on the optimization problem at hand. Common techniques include:
- Use an evaluation data set: Run the organism’s code against the data and score its performance by comparing against the known correct values. This type of evaluation is especially useful for optimizing LLM prompts and agentic code.
- Measure a desired performance metric directly: For example, we can measure the speed at which the code runs on a given input, or how optimally it solves a given problem.
- Calculate quality heuristics through code inspection: Useful heuristics can be the complexity of the code (typically simpler = better), or how well the given code generalizes. Such quality criteria can oftentimes be assessed by an LLM critique.
Sampling parents
In each iteration, parents are sampled proportional to their sampling weights.
Our sampling weight calculation is based on the formulas proposed by Darwin Gödel Machines, with a few extensions. The weight for a given candidate organism is calculated from its “fitness score” , as well as a novelty bonus based on the number of existing children as follows:
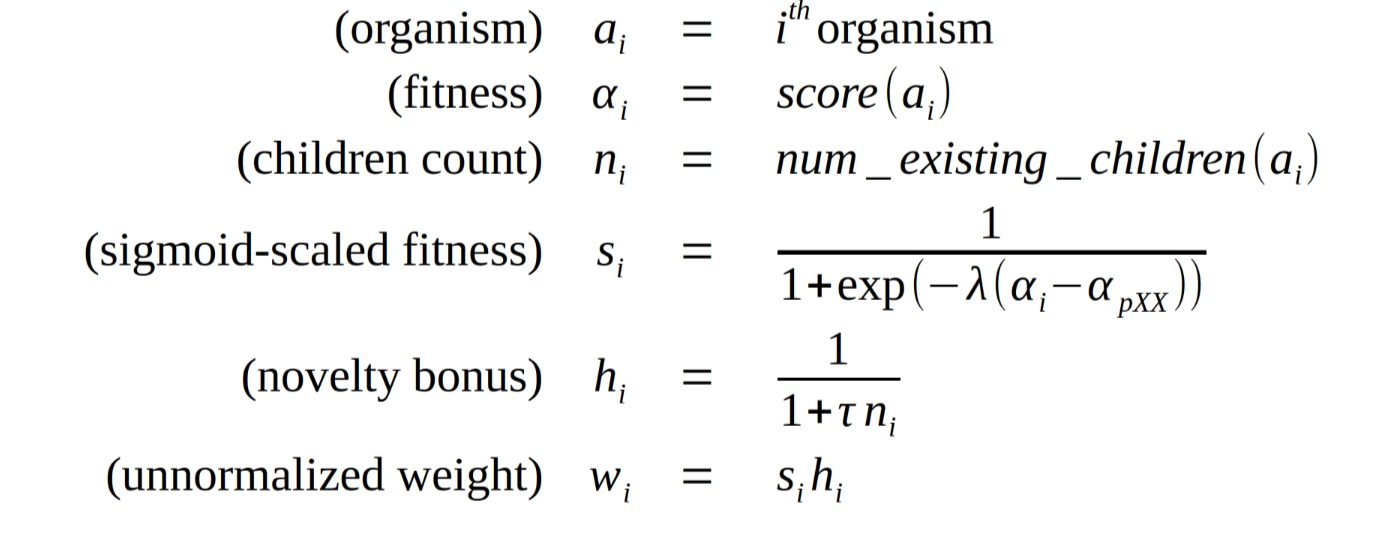
The fitness score is scaled using a sigmoid function with two hyper-parameters:
- A sharpness parameter (typical range: 5 to 20)
- and a “midpoint score”, which we calculate dynamically in each iteration to be the XXth percentile of all fitness scores currently in the population. (typical range: to )
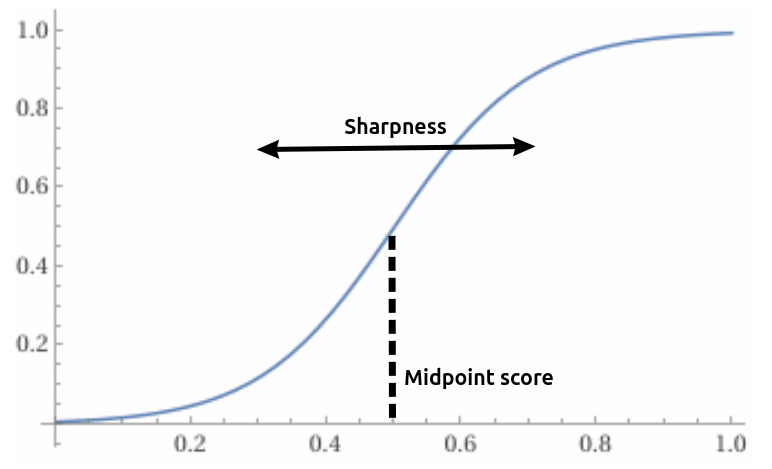
We improve upon Darwin Gödel Machines by using a dynamic, percentile-based midpoint score. By shifting the midpoint dynamically throughout the run of the evolver, we can operate in the high-gradient range of the sigmoid throughout an entire evolver run. This works even when the achievable score range is not known upfront, or when the population’s score range shifts significantly throughout the course of evolution. The dynamic midpoint score also makes it possible to use higher sharpness values in scenarios where we want to prioritize efficiency (exploitation) over diversity (exploration), without causing a premature saturation of the sigmoid-scaled scores.
The second component of the sampling weight is the novelty bonus. The novelty bonus puts emphasis on exploring newly derived organisms, and limits the amount of resources that can be spent on locally fit, but ultimately unsuccessful evolutionary “dead ends”.
We add an additional “novelty weight” hyperparameter (typically ) , which controls how quickly the novelty bonus wears off as additional children are generated for a given parent. We’ve found that certain problems benefit from a smaller novelty weight to allow for a more “thorough” exploration of high-scoring parents.
Note that the weight of any given organism is strictly positive, meaning that even organisms with low scores will be sampled occasionally. This property contributes to the evolver’s robustness in escaping local maxima, and allows it to explore a diverse range of solution approaches over time.
Mutating code
In natural evolution, mutations to DNA are random. However, making random changes to a piece of computer code and hoping that it will lead to a significant fitness improvement, or even run without error, would be extremely inefficient.
Instead, we lean on LLMs to propose targeted improvements to the parent’s code. We do not need the LLM to be reliable enough to guarantee an improvement every single time. Nor do we need it to come up with a solution that generalizes across all inputs right away. Unsuccessful improvement attempts will automatically receive a lower score, and hence become less likely to be sampled as parents further on. As long as the LLM every now and then generates a modification that leads to a better score, that solution will be picked more often moving forward and be used as a basis for further improvement.
That being said, the more often a mutation is beneficial, the more efficient the evolution process becomes. Our evolver implements several techniques to maximize the success rate of mutations, as we will discuss next.
Mutator inputs
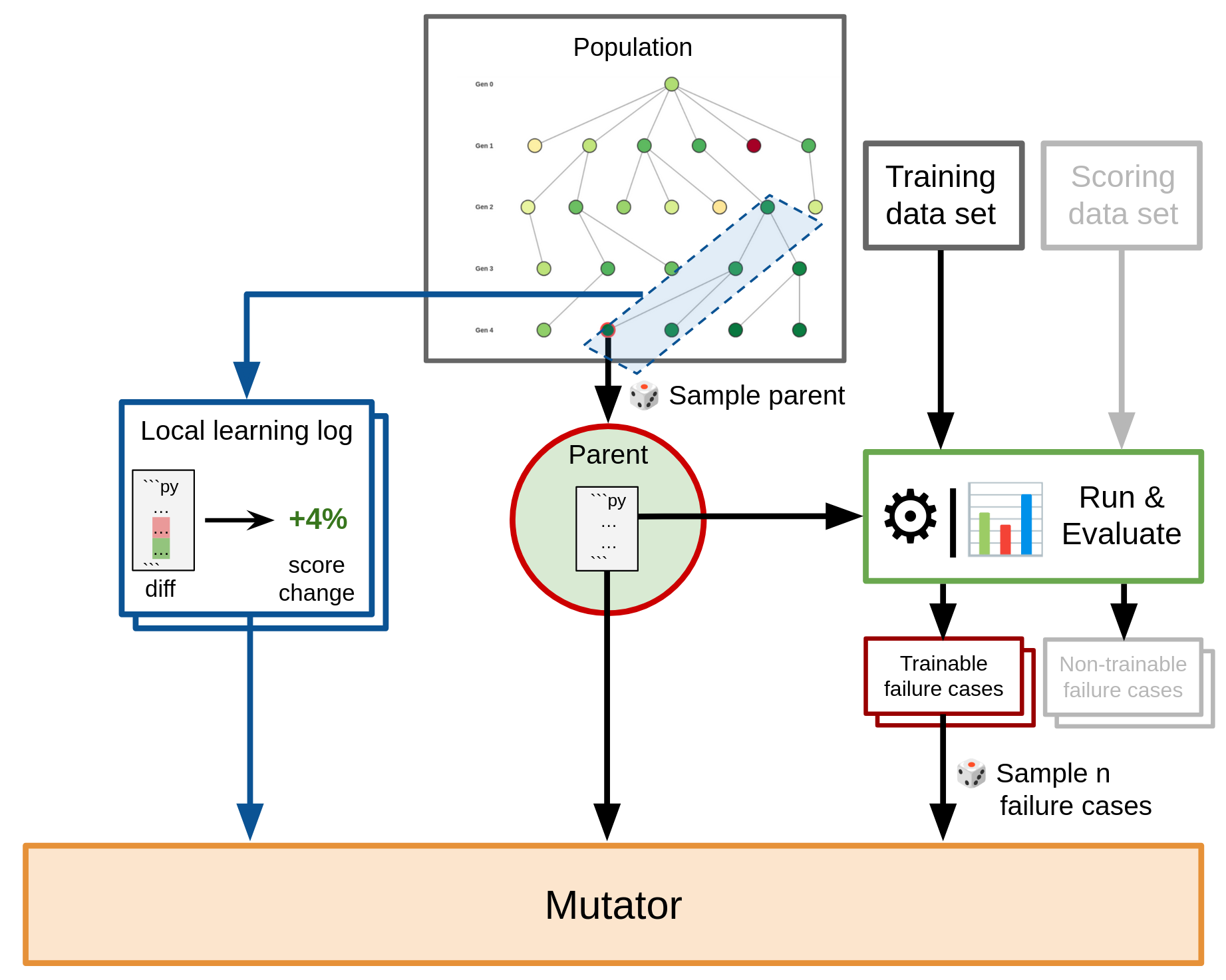
In Darwin Gödel Machines, the mutating LLM is given the parent’s code, as well as details about a single, randomly sampled, input on which the parent failed (the failure case). The failure case description for the LLM can include ground-truth information, as well as details about the parent’s performance on it (such as final outputs, debug logs, or traces). The LLM is then asked to analyze the failure case and suggest an improvement idea that is subsequently implemented.
In our Darwinian Evolver, we build upon this approach, but introduce several enhancements:
- Batch mutations: Rather than providing only a single failure case to the mutator, we support exposing and analyzing multiple failure points simultaneously. This is roughly equivalent to the use of mini-batches in Stochastic Gradient Descent. Batch mutations allow for faster progression per iteration, as a single mutator can attempt to improve over multiple failure cases in a single go.
- Separate training and scoring data sets: The training data set is used for providing mutator feedback, while a separate scoring data set is used for assigning fitness scores. This setup helps discourage mutations that are narrowly geared towards only the provided failure cases, while making sure that improvements that generalize across many inputs will be selected for.
- Learning log: The learning log is a list of past mutations together with their observed impact on the fitness score. For a given mutator call, a selection of learning log entries that come from a local neighborhood around the parent is provided. The learning log represents mutations as a code diff or change description, rather than as a full snapshot of the resulting organism. Thus, learning log entries provide a direct, differential signal on which specific change in an organism’s code led to a resulting change in performance.
- Crossover mutations: Crossover mutations are an alternative to the learning log that acts across the entire population. They work well for problems in which discoveries made in one evolutionary branch can be easily transferred into a different branch. Crossover mutations work by sampling not one, but multiple parents for a single mutation. The mutating LLM is prompted to combine the best ideas from each parent into a unified output organism. Sakana’s ShinkaEvolve also adopted a similar crossover mutator, although their variant is limited to two parents at a time. Note that while both learning log and crossover mutations can reduce overall diversity in the population, they add efficiency by allowing rare discoveries made in one ancestral line to be shared beyond the direct ancestors.
Post-mutation verification
Once the evolution process moves into a stage where further improvements are no longer trivial, we often see an increasing rate of mutator applications that do in fact not yield an improved child organism.
Such non-improvements can bog down the process: More and more similarly performing organisms need to be scored and added to the population. This is wasteful, because scoring an organism can incur significant dollar and runtime cost. Furthermore, when a rare breakthrough does eventually happen, the sheer number of lower performing organisms now in the population lowers the likelihood for the breakout organism to get sampled.
To address this problem, we introduce an optional post-mutation verification step. The post-mutation verification filters out mutations that are unlikely to provide an improvement. It is applied before scoring on the full data set is performed, potentially saving significant cost and time.
The exact shape of this post-mutation verification can differ from problem to problem. In our experiments, we have found the following strategy to be a good predictor for whether a new organism will perform better than its parent:
- Perform a “mini evaluation” on only those failure cases from its parent that were passed into the mutator.
- Dismiss the new organism if it does not show improvement on any of these failure cases.
We have frequently seen a more than 10-fold improvement in both time and cost efficiency by using this type of post-mutation verification.
What’s next?
Today, we are open-sourcing the Darwinian Evolver. Our evolver is problem-agnostic, and can be adapted to virtually any code and/or prompt optimization use case. Check out the project’s README file for examples and details on how to add your own problem specifications.
You can also download our agent verifier Vet and reap the benefits of its evolution-optimized prompts.
Follow along with our builds! Subscribe to Imbue’s email newsletter for product updates and events.
Further reading
- Out post on using the Darwinian Evolver to solve ARC-AGI tasks
- The darwinian_evolver repository on GitHub.
- Sakana.ai’s Darwin Gödel Machines and ShinkaEvolve
- Google’s AlphaEvolve
- The DSPy prompt optimizers